It is one of the most famous extinct animals of all time, ruthlessly hunted to extinction by humans in just a few decades.
Now, scientists are getting closer to returning the famous dodo to its original home in Mauritius, the island east of Madagascar in the Indian Ocean.
American startup Colossal Biosciences, based in Dallas, Texas, is using stem cell technology and genome editing to create a modern approach to the species.
Costing more than $225 million (£180 million), it is “de-extinct” the dodo more than 350 years after it was removed from Mauritius by European explorers.
And Colossal Biosciences doesn’t stop there: it also wants to recover the extinct woolly mammoth and the Tasmanian tiger.
Scientists are using stem cell technology to bring back the extinct species, more than 350 years after it went extinct.
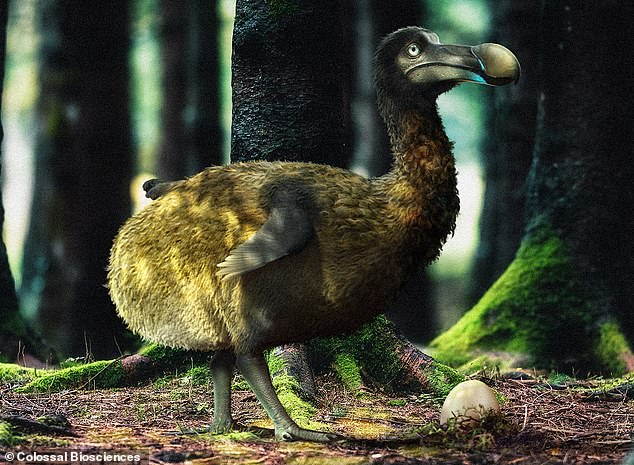

Most people believe that the dodo was a fat, gangly bird, but since it has been extinct since the late 17th century, no one knows exactly what the dodo looked like.
Colossal Biosciences has partnered with the Mauritian Wildlife Foundation to find a suitable location for the first flock of dodos once they have been grown in the laboratory.
Before being wiped out in the 17th century, this flightless bird was endemic to the island’s forests.
“The dodo, a bird intimately woven into the DNA of Mauritius, is also sadly iconic for the role humanity played in its extinction,” said Vikash Tatayah, conservation director at the Mauritian Wildlife Foundation.
‘It also symbolizes efforts to prevent species extinction.
“We are very grateful for Colossal’s technologies and the promise of returning this iconic species, extinct since the 1680s, to its native environment.”
In preparation for the release of the regenerated dodos into the wild, the two partners will collaborate on “ecosystem restoration” through invasive species removal, revegetation and community awareness efforts.
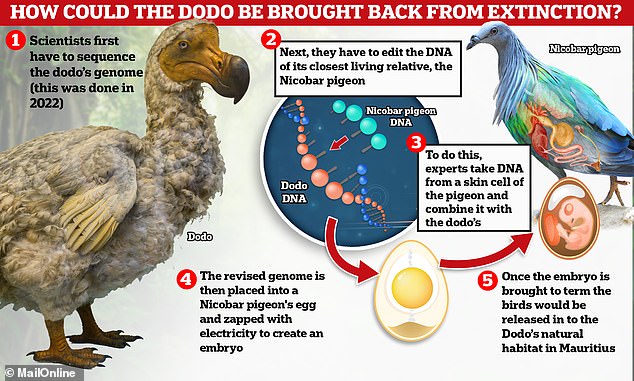
Scientists have already achieved the monumental feat of sequencing the entire genome of the extinct species, using bone samples and other fragments.
The next step is genetically editing the skin cell of a close living relative, which in the case of the dodo is the Nicobar pigeon, so that its genome matches that of the extinct bird.
The goal is to make the genome match that of the extinct bird, or come as close as possible to it, with the help of the Crispr-Cas9 gene editing tool.
This is a technique that can “cut and paste” small sections of DNA, meaning scientists can remove or edit certain genes so that they more accurately represent the animal they are seeking to recreate.
This genetically altered cell must then be used to create an embryo (just like Dolly the sheep in 1996) and carry it to term in a live surrogate mother.
Scientists hope that the chick that hatches will look like something between the Nicobar pigeon and the dodo.
Their goal is for the first laboratory-grown dodo to be born this decade.
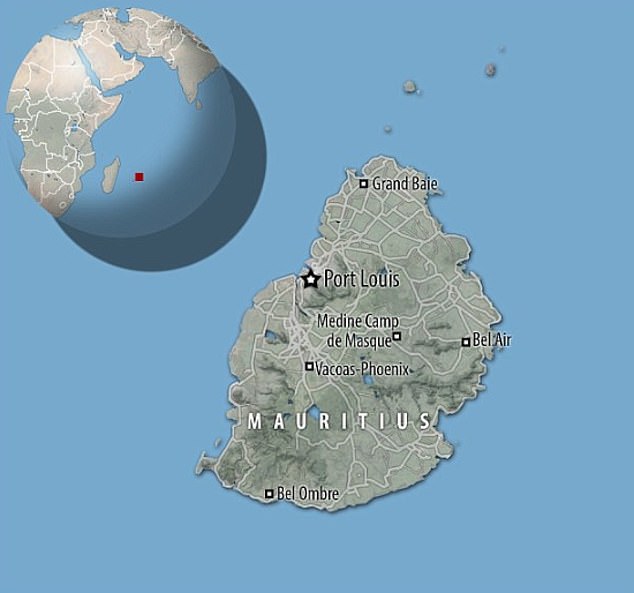
Scientists want to bring the dodo home to Mauritius, the island east of Madagascar in the Indian Ocean

The Nicobar pigeon (pictured) is the closest living relative of the dodo, found on India’s Andaman and Nicobar Islands.


The dodo gets its name from the Portuguese word for “fool,” after colonialists mocked its apparent lack of fear of human hunters.
The dodo, discovered by Europeans in 1598, was a flightless bird that lived on the island of Mauritius in the Indian Ocean.
The dodo gets its name from the Portuguese word for “fool,” after colonialists mocked its apparent lack of fear of human hunters.
He also became prey to cats, dogs and pigs that had been brought with sailors exploring the Indian Ocean.
Because the species lived in isolation in Mauritius for hundreds of years, the bird was fearless and its inability to fly made it easy prey.
Its last confirmed sighting was in 1662, after Dutch sailors first sighted the species just 64 years earlier in 1598.
Most people believe that the dodo was a fat, gangly bird, but since it has been extinct since the late 17th century, no one knows exactly what the dodo looked like.
The Oxford University Museum of Natural History is home to the only surviving remains of dodo soft tissue existing anywhere in the world.
Scientists said the ‘Oxford dodo’ was shot in the back of the head with a shotgun.

Pictured is part of the ‘Oxford dodo’, the only surviving remains of dodo soft tissue existing anywhere in the world.
There is also a dodo skull in the Zoological Museum of the University of Copenhagen and an upper jaw in the National Museum in Prague.
Now, these few fragments exist as a “symbol of human-caused extinction,” according to Colossal Biosciences.
“The dodo is a prime example of a species that became extinct because we humans made it impossible for it to survive in its native habitat,” said Professor Beth Shapiro, the company’s lead paleogeneticist.
‘Having focused on genetic advances in ancient DNA throughout my career and being the first to fully sequence the dodo genome, I am delighted to collaborate with Colossal and the people of Mauritius on the de-extinction and eventual revival of the dodo.
“In particular, I look forward to promoting genetic rescue tools focused on birds and their conservation.”
Professor Shapiro admitted that it won’t be easy to recreate a “real, living, breathing animal” in the form of a meter-tall bird.
It was his team that sequenced the bird’s entire genome for the first time in March 2022, after having spent years struggling to find sufficiently well-conserved DNA.
“Mammals are simpler,” said Professor Shapiro of the University of California, Santa Cruz.
‘If I have a cell living in a dish in the lab and I edit it to have some dodo DNA, how can I transform that cell into a real, living, breathing animal?’

However, the expert leading the dodo extinction project, paleogeneticist Beth Shapiro (pictured left), warned that it would not be easy to recreate a “real, living, breathing animal” in the form of a non-bird. flying. Ben Lamm, co-founder and CEO of Colossal is right
‘The way we can do this is to clone it, the same approach that was used to create Dolly the Sheep.
“But we don’t know how to do that with birds because of the complexities of their reproductive pathways.”
He added: “Therefore there needs to be another approach to birds and this is a really fundamental technological hurdle in de-extinction.”
“There are groups working on different approaches to achieve this and I have no doubt that we will achieve it, but it is an additional obstacle for birds that we don’t have for mammals.”
Other experts have questioned whether the new creation would really be a dodo.
“There is no one around who can teach the dodo how to be a dodo,” said Mikkel Sinding, a paleogeneticist at the University of Copenhagen. American scientist.
“You can edit the genome of something and say you’ve remade a species, but is it really the species?”